Transposable Elements (TEs) are dispersed repetitive DNA sequences in the genomes, most prevalent in eukaryotic organisms and often impacting genome evolution and architecture, mainly due to their redundancy and rearrangements they promote. For instance, TEs can comprise up to 90% in many plant genomes, e.g. maize and wheat. Conversely, independent of the number and quantity present in a given genome, TEs can also play important roles shaping gene expression and chromatin structure.
Our analysis brought an exhaustive, systematic and full-coverage genome identification in plant genomes using seven programs to annotate Transposable Elements. In both classes of TEs, several Orders and Superfamilies were found ubiquitously to all genomes. Also, 21,236,989 out of 49,802,023 TEs sequences mapped could not be classified into any of the nomenclatures known for TEs and was labeled as “Unknown” in GFF3, a standard file for annotation genes.
Many of those species do not have any identification using similar tools, as we did, to cover TEs in the species that we are providing. Although, for some plant species that TE may be quite well-annotated, we have an increase of ~71% in A. thaliana. And, for three genome-specific databases, Populus trichocarpa (black cottonwood), Glycine max (soybean) and Gossypium raimondii (cotton), we have increased the identification of TEs at 2,295%, 900% and 2,643%, respectively.
Still, the importance of studies in ncRNA:TE relationship allows us to extend and explore in some way how Transposable Elements are related to ncRNAs and vice-versa. Whilst this relationship remains unclear, we are fostering knowledge and providing a concise bulk of public data. Regarding it, our TE annotations are performed in all genomes analyzed - 67 in total - and this study has increased the knowledge base of TEs.
Here, we may use as an example A. thaliana of how our pipeline works, software dependencies, and in-house scripts we developed, which can be downloaded, used and changed freely, is available at https://github.com/daniellonghi/te_pipeline or https://github.com/alerpaschoal/apte_pipeline.
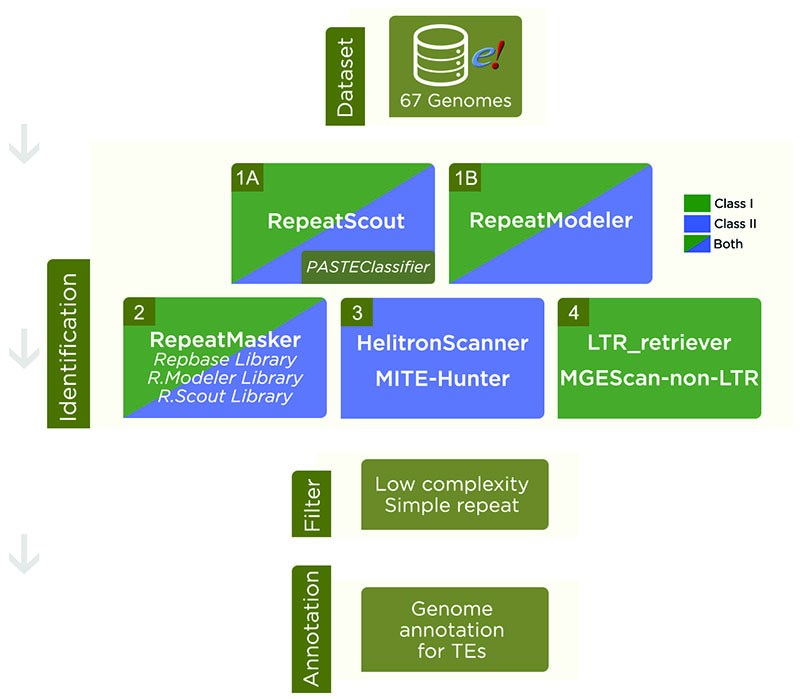
Figure 1 - Steps in Transposable Elements identification. Dataset: Genome assemblies were downloaded from Ensembl Plants. Identification: 1A) RepeatScout was used to search for putative TEs and further classification by PASTEClassifier, resulting in a library. 1B) RepeatModeler was also used to find a consensus of TEs sequences. 2) RepeatMasker was run with Repbase library and libraries from RepeatModeler and RepeatScout. 3) For Class II - Subclass 2 TEs, we also used HelitronScanner and MITE-Hunter. 4) In order to find LTR and Non-LTR retrotransposons, we used LTR_retriever and MGEScan-non-LTR, respectively. Filter: A cut-off filter was applied to remove low complexities, simple repeats and other nomenclatures that were not classified into TEs. Annotation: In result of the pipeline, we have a Transposable Element annotation for each genome analyzed.
In Download section you may able to download the annotation we deliver. If you want to access the files that we are working with by Ensembl Plants, click here, so you will be redirected to all kind of the annotation files made by them.